(In case it matters, I’m running R version 4.2.1 and brms v2.17. You can show your reader only the output from a code chunk with the nocommands option in the code fence info tag. Absolutely amazing tutorial on his blog that I tweaked only slightly. Mutate(width = CI_high - CI_low) # compute interval widthsĪll credit for the above goes to Solomon Kurz by the way. f= sim_d_and_fit_easystats, n = 50)) |> # change sample size as necessary Mutate(treatment = ifelse(group = "control", 0, 1), Prior = c(prior(normal(42.5, 5), class = Intercept), Rnorm(n, mean = mu_UNstruc, sd = sd_UNstruc))) Rnorm(n, mean = mu_struc, sd = sd_struc), Mutate(treatment = ifelse(group = "structured", 0, 1), # define the means and SD's for the idealized data If needed, select Custom from the list of the output directories on the R Markdown toolbar and specify an alternative. By default, all output files are stored in the project root directory. The R Markdown console reports on the task completion. Parameters::model_parameters("median", ci_method = "HDI", ci=.89) |>įull code for reproducability here: pacman::p_load(brms, easystats, tidyverse, janitor, flextable) P圜harm creates an HTML file with the same name as the. I get an error that says (possible class 'character' not supported'): argument is not interpreted as logical. Having trouble adapting the iterative function to use it though the command I use ( parameters::model_parameters) to extract the estimates I need can’t read the captured output when its piped from update(). Press Alt+Enter and install the missing package.So I just tried that with a single model and it does work! It produces a single model in the environment I can pull parameter estimates from. If any code line is highlighted, hover it over: you might have not the shiny package installed. Xlab = "Waiting time to next eruption (in mins)", Hist(x, breaks = bins, col = "#75AADB", border = "white", # re-executed when inputs (input$bins) changeīins <- seq(min(x), max(x), length.out = input$bins + 1) It is "reactive" and therefore should be automatically # This expression that generates a histogram is wrapped in a call # Histogram of the Old Faithful Geyser Data. # Define server logic required to draw a histogram. # Sidebar layout with input and output definitions. # Define UI for app that draws a histogram. 
The following code sample creates three content slides: Build a presentationĬreate a new *.rmd file and select Presentation when specifying its type. You can present your R markdown content in a form of presentation. 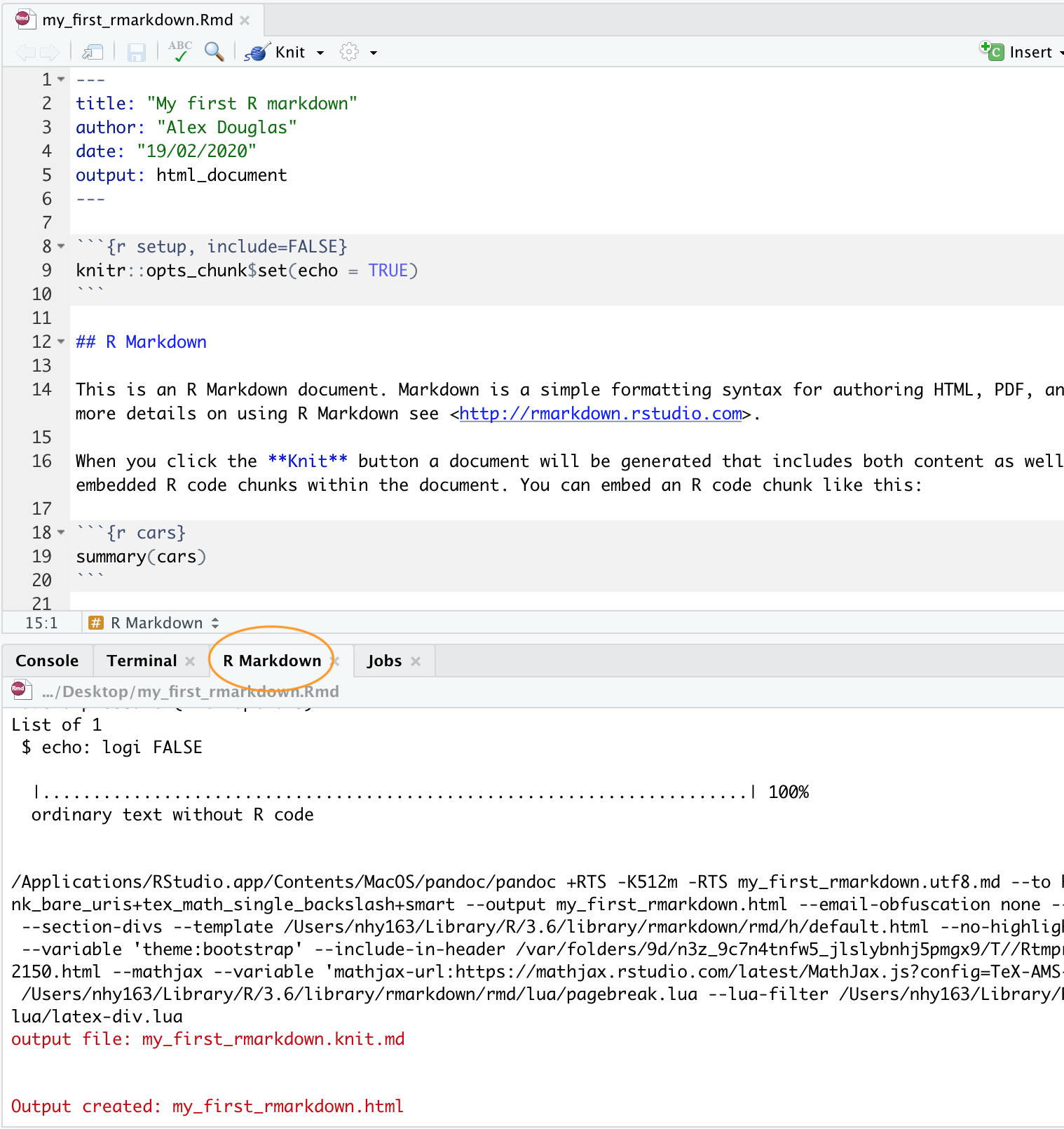
If needed, select Custom from the list of the output directories on the R Markdown toolbar and specify an alternative location for the output files.Ĭlick to open the output file in the browser. The R Markdown console reports on the task completion.īy default, all output files are stored in the project root directory. 
P圜harm creates an HTML file with the same name as the. You can create an HTML file that includes both the R Markdown source code and the results of its execution. Manage the debugging process with the debugging toolbar. The debugging process stops at a breakpoint and you can preview the current results in the Variables window. Similar to running code chunks, you cannot perform debugging, if any of the variables used in the current chunk have not been initialized. Debug code chunksīefore debugging a particular code chunk, ensure that you have debugged all chunks with its code dependencies. You can debug executable chunks in R Markdown files to detect and fix any errors in them. If needed, click to clear the execution results. You can also switch to the R Console to study variables in-detail. Preview and evaluate the results of the execution that are rendered below the chunk. For example, the second chunk in the code fragment uses the variable defined in the first chunk.Ĭlick to execute all chunks above the currently selected chunk to make sure that all required variables are initialized.Īt any time you can click to execute all remaining code chunks in the file, or opt to the execution all code chunks at once by clicking. To change the way the HTML pages are split, the splitby argument can be specified. The main difference between rendering in R Markdown and bookdown is that a book will generate multiple HTML pages by default. When executing one chunk at a time, mind code dependencies. The output format bookdown::gitbook is built upon rmarkdown::htmldocument, which was explained in Section 3.1. To execute a code chunk at the caret, click and select Run Chunk, or click on the chunk toolbar. Executes all the code chunks above the current chunk.Įxecutes all the code chunk below the current chunk.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
